One of the cool new features of KNIME Analytics Platform 3.3 is the ability to use the Java Snippet node with objects and functions that are defined in KNIME extensions. This allows an interesting and powerful new way to work with extensions that are compatible with the new functionality (more on the small amount of work required for that in a separate post, but for those who want to get a head start, here’s a link to the commit that added the functionality to the RDKit nodes). This post will demonstrate how to use the RDKit Java wrappers from within a KNIME Java Snippet node. Though this blog post uses cheminformatics functionality from the RDKit as a demonstration, the new feature can also be used to work with things like images, XML and JSON documents, and SVGs.
We’ll be working with the Java Snippet node, here’s some more information about that. Needless to say, you need to have KNIME 3.3 (or later) with the RDKit community nodes installed. As of this writing you need the nightly build of the RDKit nodes; the update site (http://update.knime.org/community-contributions/trunk) is linked from the KNIME Community site.
Example 1: Working with RDKit Molecules.
Let’s start by reading in a set of SMILES from a file, converting those to RDKit molecules, and then adding a Java Snippet node. Here’s the fragment of the workflow:
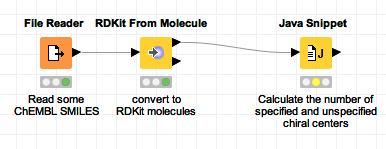
To get started, I open the empty configuration dialog of the Java Snippet node (not shown) and double click the RDKit molecule column in the column list to add a new Input:
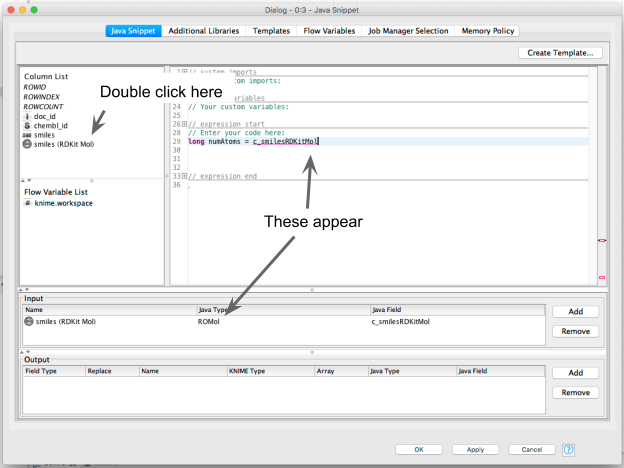
This also causes some magic to happen in the backend and the relevant dependencies to be automatically imported.
Variable completion even works, which is very cool since there’s not currently any useable documentation for the RDKit Java wrappers:
(click on the image to see it in full size)
Let’s bring in the rest of the RDKit (we could do this in a more targeted way, but this approach is simpler):
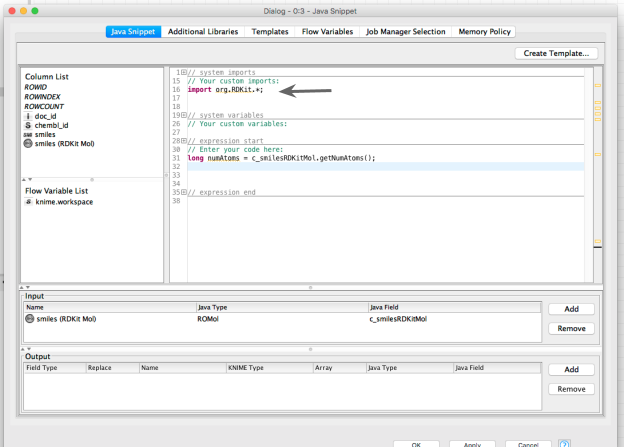
Now we can start to do the interesting stuff! Here’s a complete, but simple demonstration of using RDKit functionality to count the number of specified and unspecified chiral centers:
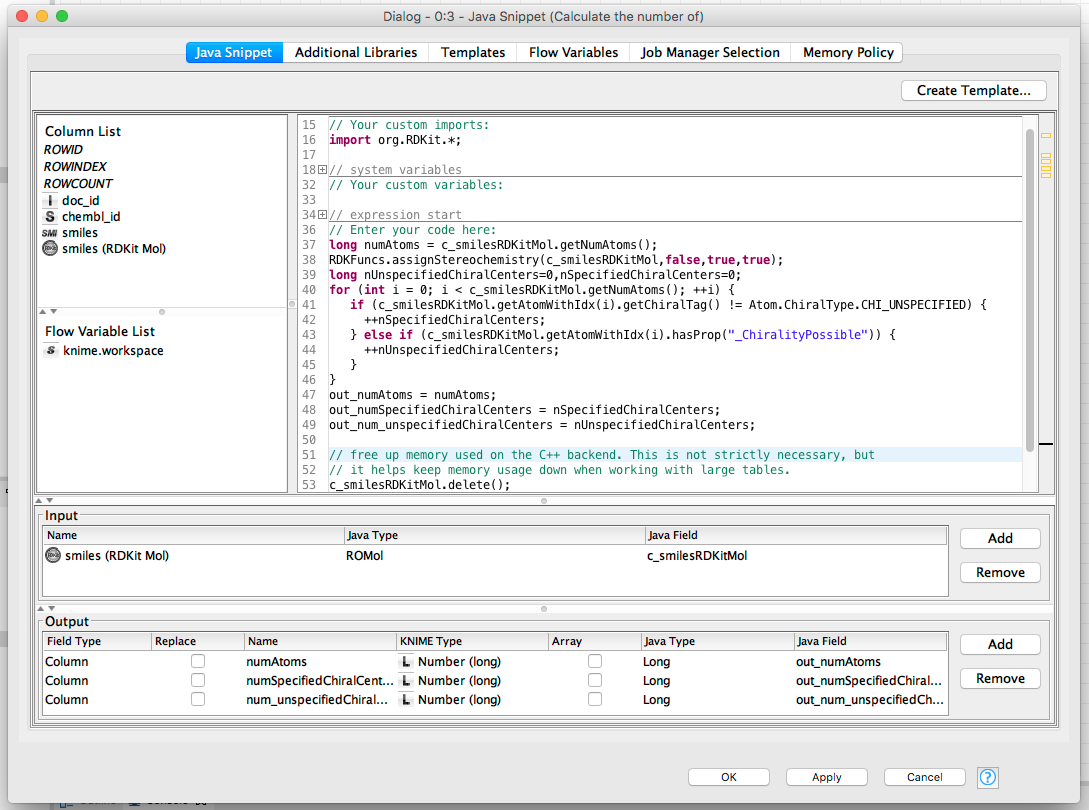
(click on the image to see it in full size)
That’s not a super-complicated example, but it already shows that you can call RDKit functions and use the methods available on the RDKit objects. The output from running the node is a new table that has three new columns added (the ones in the “Output” pane):
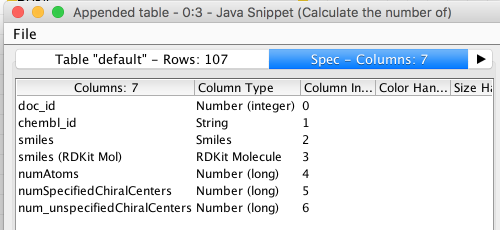
Example 2: Working with RDKit Fingerprints
Here’s another example snippet that generates a molecular fingerprint (using the RDKit’s Pattern Fingerprint function) and calculates some stats about it. The stats are returned as numbers, the fingerprint as a string of 0s and 1s:
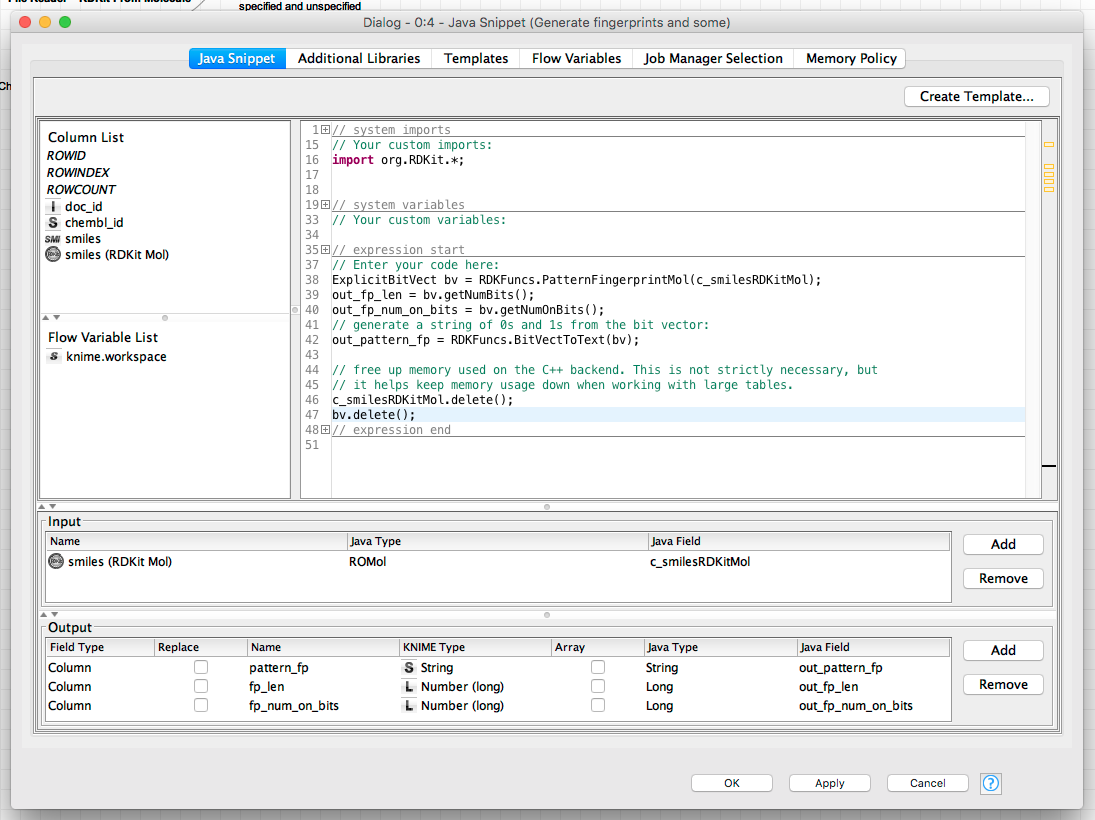
(click on the image to see it in full size)
That string of 0s and 1s can be converted into a KNIME bit vector using KNIME’s Create Bit Vector node:
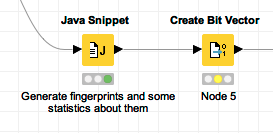
with this configuration:

Example 3: Converting KNIME bit vectors into RDKit bit vectors
For the sake of symmetry and completeness, here’s a final Java snippet that shows how to convert KNIME bit vectors, which come into the Java Snippet node as strings, into RDKit ExplicitBitVects:
(click on the image to see it in full size)
Notice that this still has the ROMol column in the inputs; this is necessary for technical reasons: without it the rest of the RDKit cannot be imported. I got that line in the inputs by double clicking the “smiles (RDKit mol)” column as in the first example snippet and then just deleting the c_smilesRDKitMol text that was inserted into the editor window. It sounds more complicated than it is!
Wrapping up
To quickly review: in this blog post I showed how KNIME’s Java Snippet node can now be used to work with custom data types like RDKit molecules and bit vectors. A future post will provide a bit more technical detail on how KNIME extension developers can add support for this to their own data types.
The workflow I created for this blog post is available on the KNIME public examples server knime://EXAMPLES/99_Community/03_RDKit/07_Java_Snippet_Exampleknime://EXAMPLES/99_Community/03_RDKit/07_Java_Snippet_Example*. You can access it directly in your version of the KNIME Analytics Platform by logging into the EXAMPLES server (in your “KNIME Explorer” pane) and browsing to the folder 99_Community/03_RDKit
Additional Resources
- KNIME Node Guide example for the Java Snippet Node: https://www.knime.org/nodeguide/scripting/java/example-of-java-snippet
- The workflow used in this blog post: knime://EXAMPLES/99_Community/03_RDKit/07_Java_Snippet_Exampleknime://EXAMPLES/99_Community/03_RDKit/07_Java_Snippet_Example*
- The github page for the RDKit KNIME nodes: https://github.com/rdkit/knime-rdkit
- The RDKit home page: http://www.rdkit.org
* The link will open the workflow directly in KNIME Analytics Platform (requirements: Windows; KNIME Analytics Platform must be installed with the Installer version 3.2.0 or higher)
